n_small <- 10
# Individual treatment effects: β_i ~ N(-0.5, 1)
# Negative on average: hospitalization helps (lower score = better health)
beta <- rnorm(n_small, mean = -0.5, sd = 1)
# Y(0): health score if NOT hospitalised (higher = worse health)
Y0 <- rnorm(n_small, mean = 3, sd = 0.5)
# Y(1): health score if hospitalised
Y1 <- Y0 + betaPotential Outcomes: From Theory to Code
Applied Causal Analysis — Coding Exercise
LMU Munich · Chair of Statistics and Econometrics
2026-04-20
Part I: The Fundamental Problem
What a researcher wishes they could see
Recall from the lecture: for each unit \(i\), we have two potential outcomes
\[Y_i = D_i \cdot Y_i(1) + (1 - D_i) \cdot Y_i(0)\]
but we only ever observe one of \(Y_i(0)\) or \(Y_i(1)\).
Let us make this concrete with a small example.
Generating the Full Schedule of Potential Outcomes
Think of the health care example. We observe \(n = 10\) patients.
Note
\(\beta_i = Y_i(1) - Y_i(0)\) is the individual treatment effect — the quantity we can never directly observe.
The Full Potential Outcomes Table
This is what a god-like researcher would see — both columns at once:
| \(i\) | \(Y_i(0)\) | \(Y_i(1)\) | \(\beta_i\) |
|---|---|---|---|
| 1 | 2.05 | 0.61 | -1.44 |
| 2 | 3.46 | 2.92 | -0.54 |
| 3 | 2.78 | 3.11 | 0.33 |
| 4 | 3.07 | 2.14 | -0.94 |
| 5 | 2.29 | 1.47 | -0.81 |
| 6 | 2.45 | -0.18 | -2.63 |
| 7 | 3.64 | 5.65 | 2.01 |
| 8 | 3.64 | 2.02 | -1.63 |
| 9 | 2.35 | 2.02 | -0.33 |
| 10 | 2.89 | 2.96 | 0.08 |
Now Assign Treatment — and Watch Data Disappear
| \(i\) | \(D_i\) | \(Y_i(0)\) | \(Y_i(1)\) | \(Y_i^{obs}\) |
|---|---|---|---|---|
| 1 | 0 | 2.05 | NA | 2.05 |
| 2 | 1 | NA | 2.92 | 2.92 |
| 3 | 0 | 2.78 | NA | 2.78 |
| 4 | 1 | NA | 2.14 | 2.14 |
| 5 | 0 | 2.29 | NA | 2.29 |
| 6 | 0 | 2.45 | NA | 2.45 |
| 7 | 0 | 3.64 | NA | 3.64 |
| 8 | 0 | 3.64 | NA | 3.64 |
| 9 | 1 | NA | 2.02 | 2.02 |
| 10 | 1 | NA | 2.96 | 2.96 |
Every ? is a missing counterfactual — this is what makes causal inference a missing data problem.
The True ATE — and Why We Can’t Compute It
[1] -0.591Warning
With only \(n = 10\) units and random assignment, these two numbers may already differ noticeably. Why? Small samples, not bias. Let us scale up.
Part II: Selection Bias
Scaling Up — and Introducing Selection
We now use \(n = 1{,}000\) units and introduce selection on levels: sicker patients (higher \(Y_i(0)\)) are more likely to seek hospitalisation.
A Biased Assignment Mechanism
Fraction treated: 0.85 Mean Y(0) among treated: 3.148 Mean Y(0) among untreated: 1.966 Important
The treated have a higher baseline \(Y_i(0)\) — they are sicker to begin with. This is exactly selection on levels.
The Bias Decomposition — in Code
Recall from the lecture:
\[\underbrace{\mathbb{E}[Y_i|D_i=1] - \mathbb{E}[Y_i|D_i=0]}_{\text{APE (observed)}} = \underbrace{\mathbb{E}[Y_i(1)-Y_i(0)|D_i=1]}_{\text{ATT}} + \underbrace{\mathbb{E}[Y_i(0)|D_i=1] - \mathbb{E}[Y_i(0)|D_i=0]}_{\text{selection bias}}\]
The Numbers
| Quantity | Value |
|---|---|
| APE (observed) | 0.678 |
| ATT | -0.504 |
| Selection Bias | 1.182 |
| ATT + Selection Bias | 0.678 |
| True ATE | -0.501 |
Note
ATT + Selection Bias = APE ✓ — the decomposition holds exactly.
The selection bias is positive: hospitalized patients are sicker to begin with, inflating the observed health gap and masking the true (negative) treatment effect.
Visualising the Selection Problem
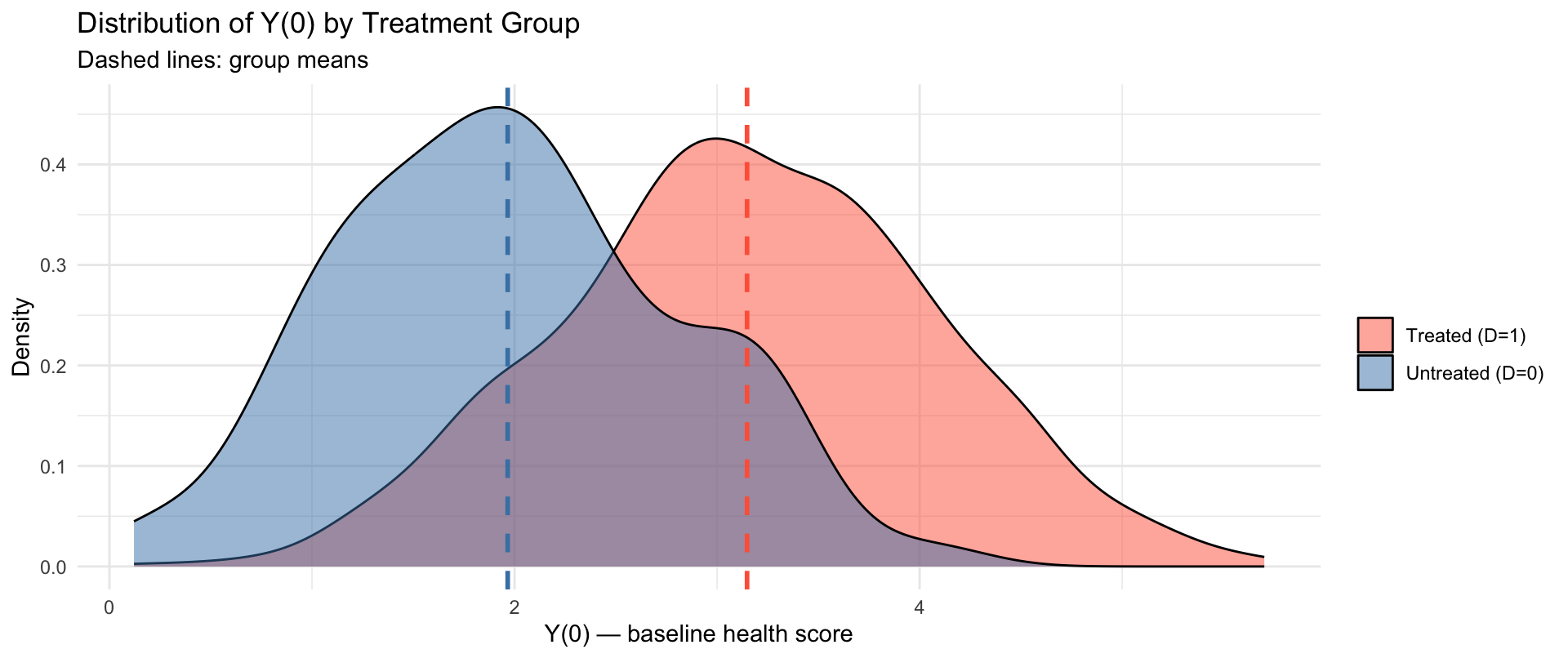
The two distributions of \(Y_i(0)\) are shifted — the treated were sicker before treatment.
Part III: Random Assignment as the Fix
Randomly Assigning Treatment
Mean Y(0) among treated: 2.969 Mean Y(0) among untreated: 2.97 Tip
Under random assignment, the two groups have similar baseline \(Y_i(0)\). Selection bias vanishes.
The Decomposition Under Random Assignment
| Quantity | Selection on Levels | Random Assignment |
|---|---|---|
| APE (observed) | 0.678 | -0.482 |
| ATT | -0.504 | -0.481 |
| ATU | NA | -0.520 |
| True ATE | -0.501 | -0.501 |
| Selection Bias | 1.182 | -0.001 |
| ATT + Selection Bias | 0.678 | -0.482 |
Side-by-Side: What Randomisation Fixes
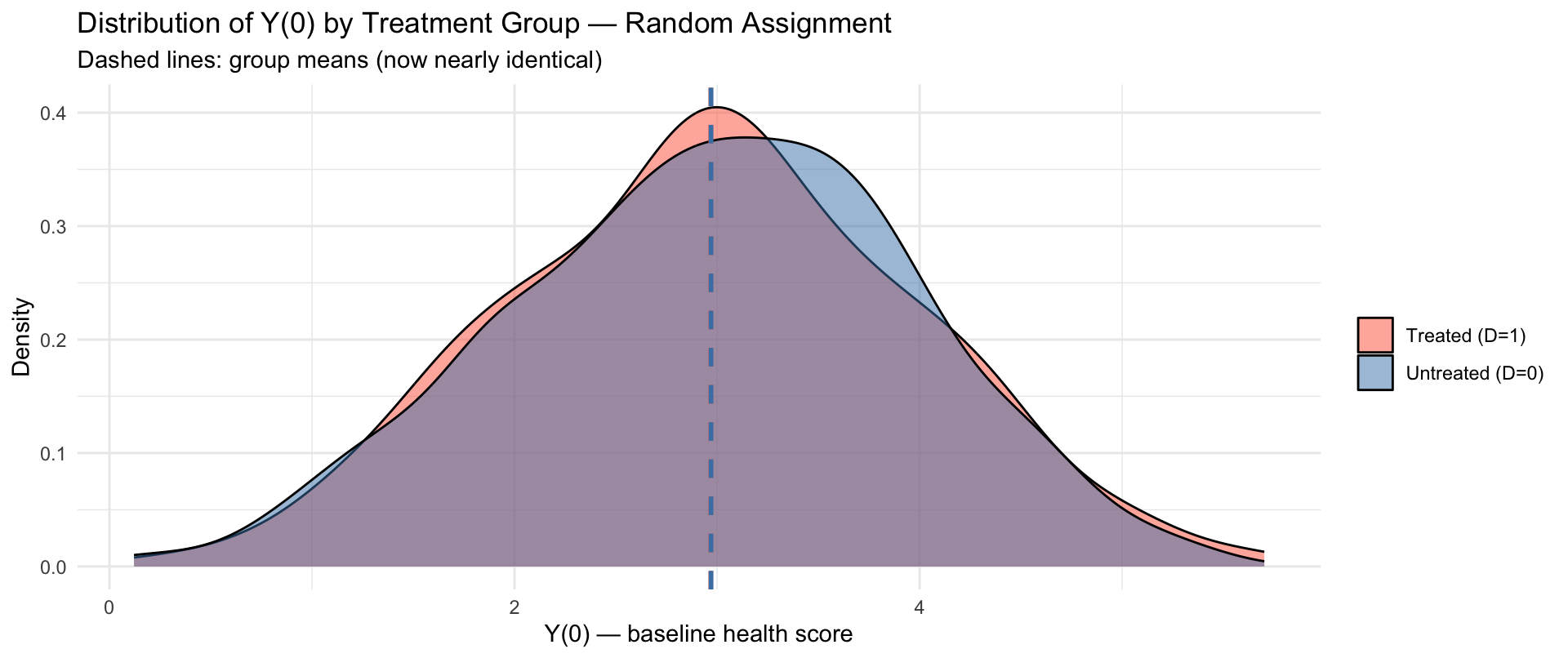
The distributions of \(Y_i(0)\) now overlap almost perfectly. Selection bias \(\approx 0\).
Randomisation Works In Expectation
Does it always work in a single draw? Let us check across many replications:
Mean of APE across 2000 replications: -0.5007 True ATE: -0.5012 Sampling Distribution of the APE
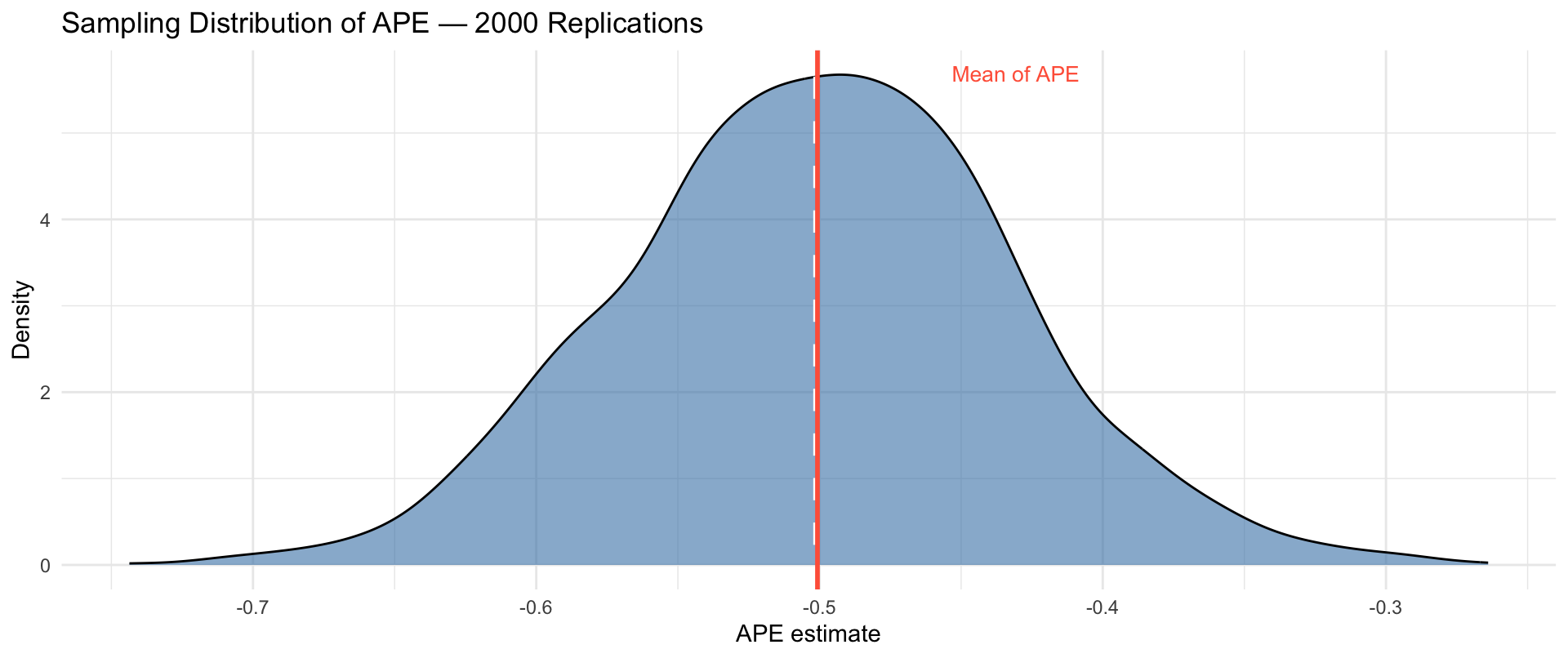
The APE is centred on the true ATE. In any single study there is sampling variability — but the estimator is unbiased.
Part IV: Selection on Gains
A Different Kind of Selection
In-class exercise (Lecture 3, Slide 19)
Show that in general \(\text{ATE} \neq \text{ATT}\) due to selection on gains, but that \(\text{ATE} = \text{ATT}\) under random assignment.
Selection on gains means individuals sort into treatment based on how much they benefit:
\[\mathbb{E}[Y_i(1) - Y_i(0) \mid D_i = 1] \neq \mathbb{E}[Y_i(1) - Y_i(0) \mid D_i = 0]\]
Generating Selection on Gains
Mean β_i among treated: -0.761 Mean β_i among untreated: 0.722 True ATE: -0.501 The treated have more negative \(\beta_i\) on average — they are self-selecting because they gain more from treatment.
ATE vs ATT vs ATU Under Selection on Gains
| Quantity | Value |
|---|---|
| True ATE | -0.501 |
| ATT | -0.761 |
| ATU | 0.722 |
| APE (observed) | -0.710 |
| Selection Bias (levels) | 0.051 |
\(\text{ATE} \neq \text{ATT} \neq \text{ATU}\) — each answers a different causal question.
Random Assignment Restores ATE = ATT
| Quantity | Selection on Gains | Random Assignment |
|---|---|---|
| ATE | -0.501 | -0.501 |
| ATT | -0.761 | -0.481 |
| ATU | 0.722 | -0.520 |
Under random assignment, \(D_i \perp (Y_i(0), Y_i(1))\), so who ends up treated is unrelated to their gains — and ATE \(\approx\) ATT \(\approx\) ATU. ✓
Part V: QTE Identification
Beyond Means — Quantile Treatment Effects
Recall from Lecture 3: under random assignment,
\[\underbrace{F_1^{-1}(\tau) - F_0^{-1}(\tau)}_{\text{QTE}_\tau,\ \text{unobserved}} = \underbrace{F^{-1}(\tau \mid D_i=1) - F^{-1}(\tau \mid D_i=0)}_{\text{observed}}\]
Let us verify this numerically.
Computing QTEs
quantiles <- seq(0.1, 0.9, by = 0.05)
# Unobserved QTE: quantiles of potential outcome distributions
qte_true <- quantile(Y1, quantiles) - quantile(Y0, quantiles)
# Observed under random assignment
qte_obs <- quantile(Y_rand[D_rand == 1], quantiles) -
quantile(Y_rand[D_rand == 0], quantiles)
# Observed under selection on levels (biased)
qte_sel <- quantile(Y_sel[D_sel == 1], quantiles) -
quantile(Y_sel[D_sel == 0], quantiles)QTE: Random Assignment Identifies the Truth
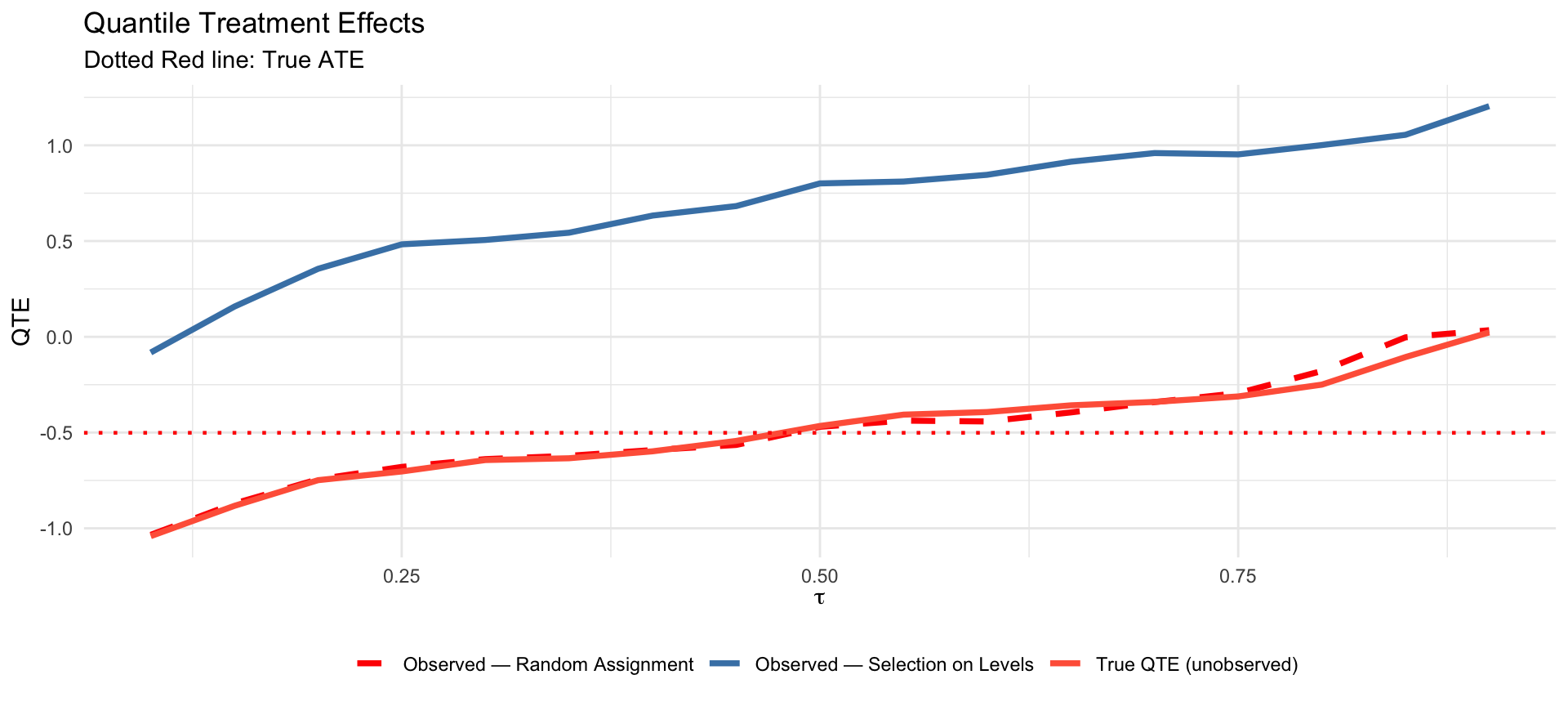
The observed QTE under random assignment (blue) tracks the true QTE closely. Under selection on levels (red) it is systematically distorted.
Summary
What We Did — and Why It Matters
The fundamental problem — with \(n=10\) we saw the missing counterfactual table directly: every
?is what makes causal inference hardSelection on levels — sicker patients self-select into hospitals; the bias decomposition \(\text{APE} = \text{ATT} + \text{Selection Bias}\) holds exactly in the data
Random assignment — the fix: \(D_i \perp (Y_i(0), Y_i(1))\) kills selection bias, and the APE is unbiased for the ATE across replications
Selection on gains — a subtler problem: ATE \(\neq\) ATT \(\neq\) ATU; random assignment equalises all three
QTE identification — under random assignment, observed quantile differences recover the true QTE; under selection they do not
Looking Ahead
The next step: estimation and inference — how do we attach uncertainty to these estimates?
- Standard errors for the difference-in-means
- Permutation tests (no asymptotic approximation needed)
- Bootstrap inference
- Covariate adjustment for efficiency gains
Reading for next class: Lecture 3 slides, sections on covariate balance and the CATE.
Applied Causal Analysis · Summer Semester 2026 · LMU Munich